REVE: A Foundation Model for EEG
Adapting to Any Setup with
Large-Scale Pretraining on 25,000 Subjects
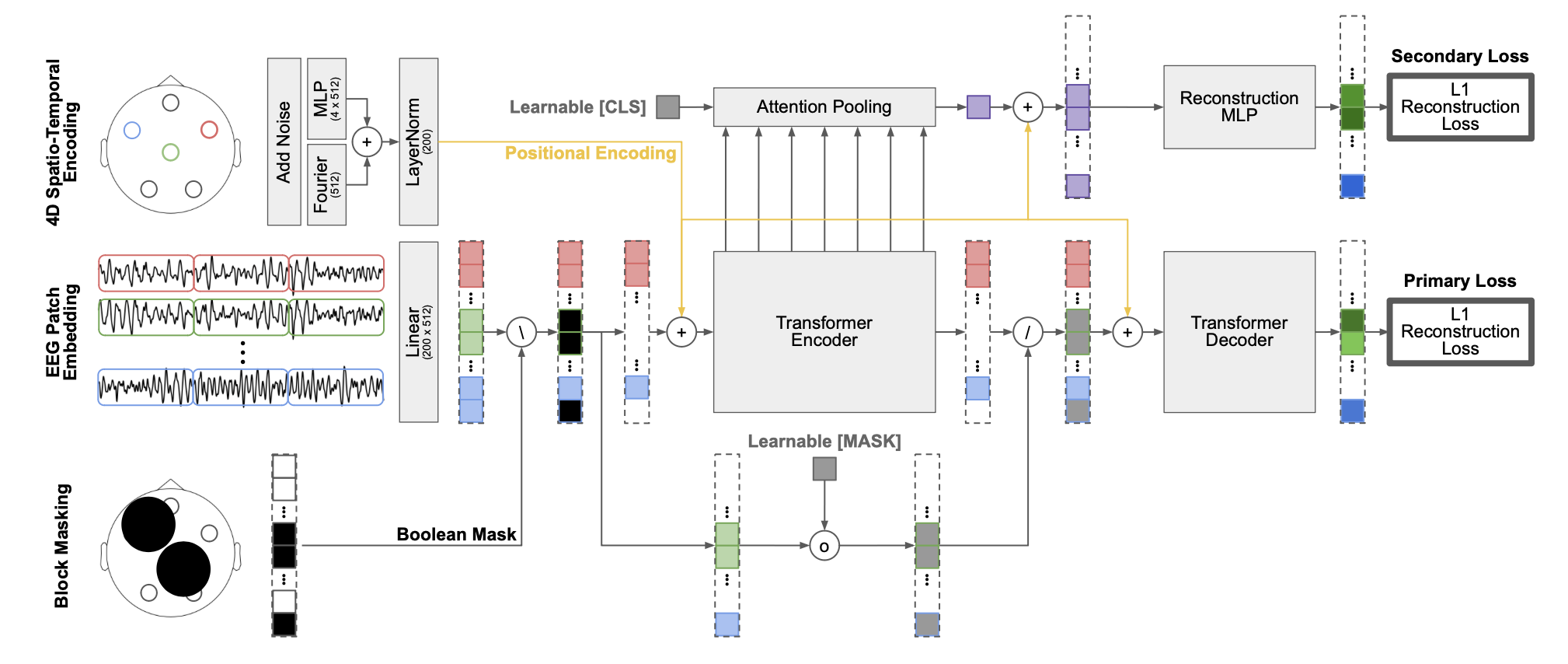
Overview of the REVE architecture and training pipeline.
Abstract
Foundation models have transformed AI by reducing reliance on task-specific data through large-scale pretraining. While successful in language and vision, their adoption in EEG has lagged due to the heterogeneity of public datasets, which are collected under varying protocols, devices, and electrode configurations. Existing EEG foundation models struggle to generalize across these variations, often restricting pretraining to a single setup, resulting in suboptimal performance, in particular under linear probing. We present REVE (Representation for EEG with Versatile Embeddings), a pretrained model explicitly designed to generalize across diverse EEG signals. REVE introduces a novel 4D positional encoding scheme that enables it to process signals of arbitrary length and electrode arrangement. Using a masked autoencoding objective, we pretrain REVE on over 60,000 hours of EEG data from 92 datasets spanning 25,000 subjects, representing the largest EEG pretraining effort to date. REVE achieves state-of-the-art results on 10 downstream EEG tasks, including motor imagery classification, seizure detection, sleep staging, cognitive load estimation, and emotion recognition. With little to no fine-tuning, it demonstrates strong generalization, and nuanced spatio-temporal modeling.
Changelog
March 2026
Code and dataset release
We release the code to pretrain/fine-tune REVE and
the
open part of the pretraining dataset.
December 2025
NeurIPS and Braindecode
We present REVE at NeurIPS 2025 and integrate the
model
into Braindecode v1.3.0.
October 2025
Model release
REVE is now available as a pretrained model on Hugging
Face. We also release a Colab tutorial to fine-tune REVE on EEGMAT.
September 2025
Announcement
REVE accepted as a poster at NeurIPS 2025.
We release the code to pretrain/fine-tune REVE and the open part of the pretraining dataset.
We present REVE at NeurIPS 2025 and integrate the model into Braindecode v1.3.0.
REVE is now available as a pretrained model on Hugging Face. We also release a Colab tutorial to fine-tune REVE on EEGMAT.
REVE accepted as a poster at NeurIPS 2025.
Code example
from transformers import AutoModel
pos_bank = AutoModel.from_pretrained("brain-bzh/reve-positions", trust_remote_code=True)
model = AutoModel.from_pretrained("brain-bzh/reve-base", trust_remote_code=True)
eeg_data = ... # EEG data (batch_size, channels, time_points), must be sampled at 200 Hz
electrode_names = [...] # List of electrode names corresponding to the channels in eeg_data
positions = pos_bank(electrode_names) # Get positions (channels, 3)
positions = positions.expand(eeg_data.size(0), -1, -1) # Expand to (batch_size, channels, 3)
output = model(eeg_data, positions)
Dataset Composition (Proportion of Total Hours)
Data Sources
Channel Counts
Tables
Main results (1/2)
| Methods | TUAB | TUEV | PhysioNetMI | BCI-IV-2a | FACED |
| EEGNet | 0.7642 ± 0.0036 | 0.3876 ± 0.0143 | 0.5814 ± 0.0125 | 0.4482 ± 0.0094 | 0.4090 ± 0.0122 |
| EEGConformer | 0.7758 ± 0.0049 | 0.4074 ± 0.0164 | 0.6049 ± 0.0104 | 0.4696 ± 0.0106 | 0.4559 ± 0.0125 |
| SPaRCNet | 0.7896 ± 0.0018 | 0.4161 ± 0.0262 | 0.5932 ± 0.0152 | 0.4635 ± 0.0117 | 0.4673 ± 0.0155 |
| ContraWR | 0.7746 ± 0.0041 | 0.4384 ± 0.0349 | 0.5892 ± 0.0133 | 0.4678 ± 0.0125 | 0.4887 ± 0.0078 |
| CNN-Transformer | 0.7777 ± 0.0022 | 0.4087 ± 0.0161 | 0.6053 ± 0.0118 | 0.4600 ± 0.0108 | 0.4697 ± 0.0132 |
| FFCL | 0.7848 ± 0.0038 | 0.3979 ± 0.0104 | 0.5726 ± 0.0092 | 0.4470 ± 0.0143 | 0.4673 ± 0.0158 |
| ST-Transformer | 0.7966 ± 0.0023 | 0.3984 ± 0.0228 | 0.6035 ± 0.0081 | 0.4575 ± 0.0145 | 0.4810 ± 0.0079 |
| BIOT | 0.7959 ± 0.0057 | 0.5281 ± 0.0225 | 0.6153 ± 0.0154 | 0.4748 ± 0.0093 | 0.5118 ± 0.0118 |
| LaBraM-Base | 0.8140 ± 0.0019 | 0.6409 ± 0.0065 | 0.6173 ± 0.0122 | 0.4869 ± 0.0085 | 0.5273 ± 0.0107 |
| CbraMod | 0.8289 ± 0.0022 | 0.6671 ± 0.0107 | 0.6417 ± 0.0091 | 0.5138 ± 0.0066 | 0.5509 ± 0.0089 |
| REVE-Base | 0.8315 ± 0.0014 | 0.6759 ± 0.0229 | 0.6480 ± 0.0140 | 0.6396 ± 0.0095 | 0.5646 ± 0.0164 |
Main results (2/2)
| Methods | ISRUC | Mumtaz | MAT | BCI-2020-3 | Average |
| EEGNet | 0.7154 ± 0.0121 | 0.9232 ± 0.0104 | 0.6770 ± 0.0116 | 0.4413 ± 0.0096 | 0.5941 ± 0.0037 |
| EEGConformer | 0.7400 ± 0.0133 | 0.9308 ± 0.0117 | 0.6805 ± 0.0123 | 0.4506 ± 0.0133 | 0.6128 ± 0.0044 |
| SPaRCNet | 0.7487 ± 0.0075 | 0.9316 ± 0.0095 | 0.6879 ± 0.0107 | 0.4426 ± 0.0156 | 0.6156 ± 0.0047 |
| ContraWR | 0.7402 ± 0.0126 | 0.9195 ± 0.0115 | 0.6631 ± 0.0097 | 0.4257 ± 0.0162 | 0.6119 ± 0.0053 |
| CNN-Transformer | 0.7363 ± 0.0087 | 0.9305 ± 0.0068 | 0.6779 ± 0.0268 | 0.4533 ± 0.0092 | 0.6133 ± 0.0045 |
| FFCL | 0.7277 ± 0.0182 | 0.9314 ± 0.0038 | 0.6798 ± 0.0142 | 0.4678 ± 0.0197 | 0.6085± 0.0044 |
| ST-Transformer | 0.7381 ± 0.0205 | 0.9135 ± 0.0103 | 0.6631 ± 0.0173 | 0.4126 ± 0.0122 | 0.6071 ± 0.0048 |
| BIOT | 0.7527 ± 0.0121 | 0.9358 ± 0.0052 | 0.6875 ± 0.0186 | 0.4920 ± 0.0086 | 0.6438 ± 0.0044 |
| LaBraM-Base | 0.7633 ± 0.0102 | 0.9409 ± 0.0079 | 0.6909 ± 0.0125 | 0.5060 ± 0.0155 | 0.6653 ± 0.0031 |
| CBraMod | 0.7865 ± 0.0110 | 0.9560 ± 0.0056 | 0.7256 ± 0.0132 | 0.5373 ± 0.0108 | 0.6898 ± 0.0031 |
| REVE-Base | 0.7819 ± 0.0078 | 0.9644 ± 0.0097 | 0.7660 ± 0.0355 | 0.5635 ± 0.0123 | 0.7150 ± 0.0057 |
Linear probing performance against CBraMod
| Dataset | REVE-B |
REVE-B | REVE-L |
REVE-L | CBraMod |
CBraMod |
| Mumtaz | 0.962 ± 0.003 | 0.931 ± 0.021 | 0.985 ± 0.006 | 0.980 ± 0.009 | 0.859 ± 0.009 | 0.907 ± 0.027 |
| M. Arithmetic | 0.725 ± 0.010 | 0.740 ± 0.073 | 0.712 ± 0.008 | 0.665 ± 0.103 | 0.500 ± 0.000 | 0.605 ± 0.020 |
| TUAB | 0.810 ± 0.007 | 0.809 ± 0.004 | 0.821 ± 0.004 | 0.809 ± 0.004 | 0.500 ± 0.000 | 0.500 ± 0.000 |
| PhysioNetMI | 0.537 ± 0.005 | 0.510 ± 0.012 | 0.551 ± 0.001 | 0.617 ± 0.000 | 0.256 ± 0.002 | 0.531 ± 0.015 |
| BCIC-IV-2a | 0.432 ± 0.004 | 0.517 ± 0.015 | 0.534 ± 0.001 | 0.603 ± 0.011 | 0.287 ± 0.023 | 0.376 ± 0.006 |
| ISRUC | 0.697 ± 0.011 | 0.662 ± 0.030 | 0.743 ± 0.004 | 0.758 ± 0.001 | 0.407 ± 0.049 | 0.430 ± 0.043 |
| HMC | 0.647 ± 0.008 | 0.604 ± 0.008 | 0.703 ± 0.003 | 0.710 ± 0.007 | 0.368 ± 0.001 | 0.538 ± 0.009 |
| BCIC2020-3 | 0.234 ± 0.009 | 0.390 ± 0.017 | 0.274 ± 0.001 | 0.378 ± 0.021 | 0.214 ± 0.003 | 0.374 ± 0.007 |
| TUEV | 0.592 ± 0.008 | 0.508 ± 0.073 | 0.630 ± 0.003 | 0.550 ± 0.014 | 0.219 ± 0.009 | 0.482 ± 0.037 |
| Faced | 0.240 ± 0.010 | 0.422 ± 0.028 | 0.283 ± 0.003 | 0.469 ± 0.007 | 0.117 ± 0.005 | 0.261 ± 0.013 |
| Average | 0.586 | 0.609 | 0.623 | 0.654 | 0.373 | 0.501 |
Electrode Positions
The diverse set of electrodes used in REVE's pretraining.
BibTeX
@article{elouahidi2025reve,
title = {{REVE}: A Foundation Model for {EEG}: Adapting to Any Setup with Large-Scale Pretraining on 25,000 Subjects},
author = {El Ouahidi, Yassine and Lys, Jonathan and Thölke, Philipp and Farrugia, Nicolas and Pasdeloup, Bastien and Gripon, Vincent and Jerbi, Karim and Lioi, Giulia},
journal = {Advances in Neural Information Processing Systems},
year = {2025},
url = {https://brain-bzh.github.io/reve/}
}